Lifestyles of the fast and slow (bacteria): In the wild, most live in the slow lane
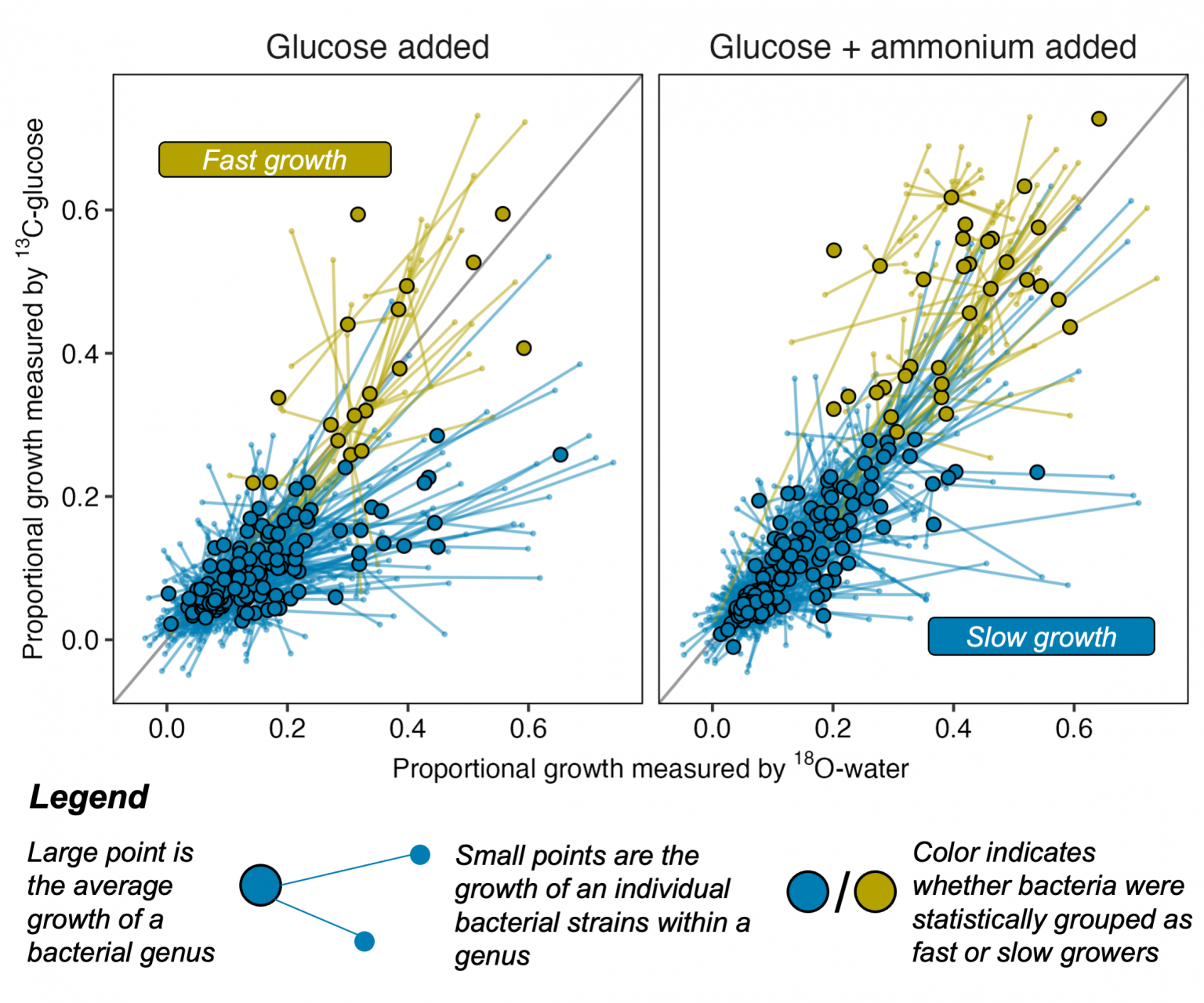
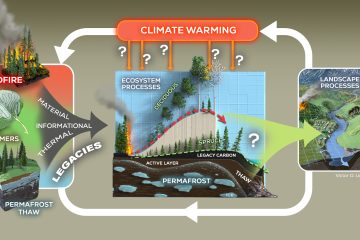
A study led by Northern Arizona University offers new evidence that a common framework to sort bacteria into two lifestyles doesn’t easily apply to bacteria living in wild soil. The findings, published in The ISME Journal, show that rather than bacteria falling into two major lifestyle groups—one adapted to be competitive and fast-growing, the other slow-growing and resistant to starvation—most bacteria observed in the wild were slow growers, with fast growers isolated to a small number of species.
“What happens in the lab and what happens in wild soil are often worlds apart, and we need to be testing and challenging ideas about bacteria and microbes from the lab with what we see in the field,” said lead author Bram Stone, who conducted the research at NAU’s Center for Ecosystem Science and Society (Ecoss) and is now a Linus Pauling Postdoctoral Fellow at Pacific Northern National Laboratory. “Many of our society’s most urgent questions about carbon storage and how soils will respond to climate change rely on understanding better how microbes act in nature.”
Suppose it is true that societal needs can sometimes accelerate the rate science is done through funding and policy prioritization. In that case, it’s also true that some fields are rapidly advancing yet still playing catchup to accelerating challenges like the climate crisis. Microbial ecology has grown by leaps and bounds in recent years as modern sequencing technology improves, becomes more widely available and gets applied in new ways. And yet, as global carbon budgets have tightened and human-caused emissions continue to fuel non-linear climate impacts, the need for knowing what the microbes will do in a warmer world has arguably accelerated even faster. Figuring out not only which microbes are where but who’s growing, who’s dying, what environmental factors affect their lives and how they interact is still a game of catchup. New data are needed to confirm or complicate some of the broad frameworks that scientists have used to make sense of this invisible world. Such conceptual frameworks and new data to test them are both necessary parts of the process to better understand microbial communities and their importance in supporting healthy soil and cycling carbon and other nutrients.
Stone says the team’s findings echo the shift from hard categories toward statistically derived trait spectrums in other fields, including psychology. (Think the move away from the Myers-Briggs test toward the trait-based spectrum of “the Big Five” personality factors.)
“Our goal is to identify the most salient microbial traits that determine actual behavior in the soil and that determine things like energy flow,” Stone said. “And we want to express those traits numerically. With a tool like that, we can make better predictions of how microbial communities react to climate change, or pollution, or a new crop rotation in an agricultural field.”
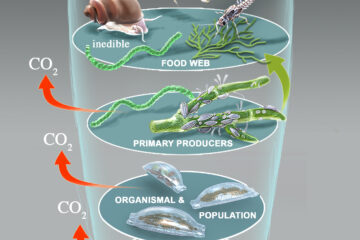
The study relies on data gathered via quantitative stable isotope probing, or qSIP, a technique that uses stable isotopes or atoms labeled with an extra neutron to track the fate of a water or sugar molecule through soil. Researchers analyze a sample of wild soil treated with this labeled water or sugar and look for where that molecular hashtag appears in DNA—meaning it has been incorporated by a microbe. By sequencing the DNA in that soil sample at different points in time, researchers at NAU, where the technique was developed, can see which microbes grew—and by how much—and how quickly the community changed.
“It’s so exciting to me that we can get the data in nature, rather than speculate,” said Bruce Hungate, director of Ecoss and a co-author of the new study. “Being able to conduct microbiology in the field like this means we can reasonably scale up to predict fluxes for an entire ecosystem or region, all while retaining the high taxonomic resolution available from modern sequencing.”
Other collaborators on the study include Paul Dijkstra, Raina Fitzpatrick, Megan Foley, Michaela Hayer, Ben Koch, Junhui Li, Ayla Martinez, Jane Marks, Rebecca Mau, Egbert Schwartz and Alicia Purcell of Ecoss, Lawrence Livermore National Laboratory, Pacific Northwest National Laboratory, University of California-Irvine, West Virginia University and the Institute for Environmental Genomics at the University of Oklahoma. This research was supported by grants from the U.S. Department of Energy’s Biological Systems Science Division Program in Genomic Science and the LLNL ‘Microbes Persist’ Soil Microbiome Scientific Focus Area and by the National Science Foundation.