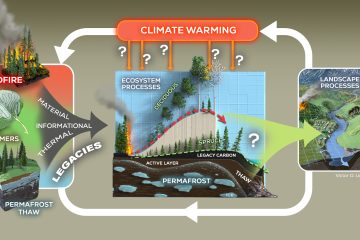
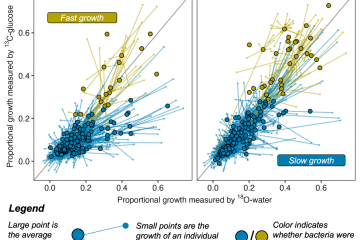
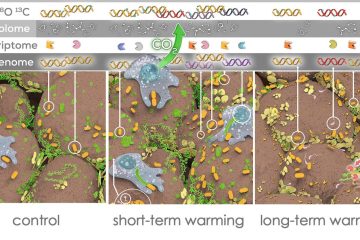
Position-specific metabolic probing and metagenomics of microbial communities reveal conserved central carbon metabolic network activities at high temperatures
Temperature is a primary driver of microbial community composition and taxonomic diversity; however, it is unclear to what extent temperature affects characteristics of central carbon metabolic pathways (CCMPs). In this study, 16S rRNA gene amplicon and metagenome sequencing were combined with 13C-labeled metabolite probing of the CCMPs to assess community carbon metabolism along a temperature gradient (60-95 °C) in Great Boiling Spring, NV. 16S rRNA gene amplicon diversity was inversely proportional to temperature, and Archaea were dominant at higher temperatures. Metagenomes spanning the temperature gradient hosted abundant CCMPs genes and many individual metagenome-assembled genomes had complete pathways. In contrast, genes encoding cellulosomes and some others involved in plant matter degradation and most for photosynthesis and were absent at higher temperatures. In situ 13C-CO2 production from labeled isotopomer pairs of glucose, pyruvate, and acetate suggested lower relative oxidative pentose phosphate pathway activity and/or fermentation at 60°C, and a stable or decreased maintenance energy demand at higher temperatures. Catabolism of 13C-labeled citrate, succinate, L-alanine, L-serine, and L-cysteine was observed at 85°C, demonstrating broad heterotrophic activity. Together, these results suggest that temperature-driven losses in biodiversity in geothermal systems may not alter CCMP function or maintenance energy demands at a community level.