Microbial rRNA synthesis and growth compared through quantitative stable isotope probing with H218O
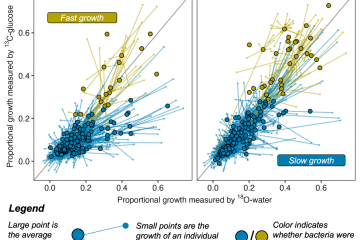
Growing bacteria have a high concentration of ribosomes to ensure sufficient protein synthesis, which is necessary for genome replication and cellular division. To elucidate whether metabolic activity of soil microorganisms is coupled with growth, we investigated the relationship between rRNA and DNA synthesis in a soil bacterial community using quantitative stable isotope probing (qSIP) with H2 18O. Most soil bacterial taxa were metabolically active and grew, and there was no significant difference between the isotopic composition of DNA and RNA extracted from soil incubated with H2 18O. The positive correlation between 18O content of DNA and rRNA of taxa, with a slope statistically indistinguishable from 1 (slope = 0.96; 95% confidence interval [CI], 0.90 to 1.02), indicated that few taxa made new rRNA without synthesizing new DNA. There was no correlation between rRNA-to-DNA ratios obtained from sequencing libraries and the atom percent excess (APE) 18O values of DNA or rRNA, suggesting that the ratio of rRNA to DNA is a poor indicator of microbial growth or rRNA synthesis. Our results support the notion that metabolic activity is strongly coupled to cellular division and suggest that nondividing taxa do not dominate soil metabolic activity. IMPORTANCE Using quantitative stable isotope probing of microbial RNA and DNA with H2 18O, we show that most soil taxa are metabolically active and grow because their nucleic acids are significantly labeled with 18O. A majority of the populations that make new rRNA also grow, which argues against the common paradigm that most soil taxa are dormant. Additionally, our results indicate that relative sequence abundance-based RNA-to-DNA ratios, which are frequently used for identifying active microbial populations in the environment, underestimate the number of metabolically active taxa within soil microbial communities.